Red tides are becoming more frequent due to climate change. Cyanobacteria and dinoflagellates found in red tides can produce saxitoxin (STX), a neurotoxin that is so poisonous that it is the only marine toxin to have been declared a chemical weapon. Shellfish can store STX for weeks to years after a red tide, and people who eat contaminated shellfish can become sick and even die from paralytic shellfish poisoning, a deadly condition that is becoming more common due to the afore mentioned climate change. STX exerts its toxic effects by blocking voltage-gated sodium channels found in neurons. Certain animals, particularly some types of frogs, are resistant to being poisoned by STX. This resistance is thought to be due to the animals’ ability to produce saxiphilin (Sxph), a protein that acts as a “toxin sponge” because it can bind to and sequester STX so that it can no longer block sodium channels. A new study, which involved experiments performed at the U.S. Department of Energy’s Advanced Photon Source (APS), shows how a particular Sxph protein, RcSxph, binds to STX and blocks its toxic effects. These findings, published in the Proceedings of the National Academy of Sciences of the United States of America, could lead to the design of biological tools that can detect or neutralize STX and related toxins, an important step in combating paralytic shellfish poisoning.
This new study is the first to define a “binding code” for how Sxph proteins bind to STX to serve their toxin sponge function. The researchers first focused on RcSxph, which is produced by the American bullfrog. They developed a set of tests that show how tightly RcSxph binds to STX. They then used these tests to see how specific mutations in the RcSxph protein changed its binding to STX. The team discovered that mutations in two binding “hot spots” affected how tightly RcSxph sequestered STX, suggesting that these spots are an important part of the binding code.
The team also identified a site that enhanced binding to STX when it was mutated. Using x-ray diffraction data collected at the National Institute of General Medical Sciences and National Cancer Institute (GM/CA-XSD) structural biology facility 23-ID-B x-ray beamline at the APS, a Department of Energy user facility at Argonne National Laboratory, the team was able to create three-dimensional crystal structures of this mutated RcSxph protein and the mutated RcSxph protein bound to STX, providing further detail about how the mutation enhanced STX binding. This information could be used to engineer more effective toxin sponges in the future.
The researchers also looked at how different mutations influenced RcSxph’s ability to act as a toxin sponge by seeing how well STX was able to block sodium channel function in the presence of the different RcSxph mutants. As expected, RcSxph proteins with mutations in the two binding hot spots were less able to stop STX from blocking the sodium channels, whereas the mutants with tighter STX binding served as more effective toxin sponges As expected, RcSxph proteins with mutations in the two binding hot spots were less able to stop STX from blocking the sodium channels, whereas the mutants with tighter STX binding served as more effective toxin sponges.
Before this study, Sxph proteins had only been identified in two other species, despite the fact that several poison-dart frog species are resistant to STX. Using their STX-binding code, the research team was able to identify 10 new Spxh proteins in frog and toad species. Surprisingly, some of these species have been separated by ~140 million years of evolution and are not known to encounter STX, and yet all produce similar toxin sponges.
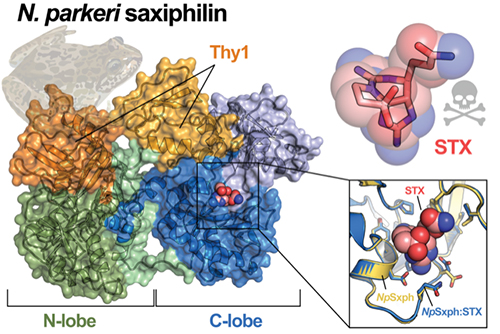
When the researchers tested four of these newly discovered proteins, they found that all four could bind to STX and two showed tighter binding than RcSxph, including one, NpSxph, from the High Himalaya frog for which they determined structures alone and bound to STX (Fig. 1).
These results show that nature follows the STX binding rules uncovered by the team and that the ability of NpSxph to sequester STX tightly comes from an amino acid change at the site identified as an affinity-enhancer in the RcSxph studies.
By identifying Sxph’s STX-binding code and determining how particular mutations can enhance binding, these findings could lead to the design of STX sensors as well as treatments for STX poisoning, a welcome possibility for shellfish lovers. ― Summer Allen
See: Zhou Chen1, Sandra Zakrzewska1, Holly S. Hajare2, Aurora Alvarez-Buylla2, Fayal Abderemane-Ali1, Maximiliana Bogan2 , Dave Ramirez2, Lauren A. O’Connell2 , J. Du Bois2 , and Daniel L. Minor, Jr.1,3, “Definition of a saxitoxin (STX) binding code enables discovery and characterization of the anuran saxiphilin family,” Proc. Natl. Acad. Sci. U.S.A. 119(44), e2210114119 (2022). DOI: 10.1073/pnas.2210114119
Author affiliations: 1University of California, San Francisco, 2Stanford University, 3Lawrence Berkeley National Laboratory
Correspondence: * daniel.minor@ucsf.edu
This work was supported by grants Department of Defense (DoD) HDTRA-1-19-1-0040 and HDTRA-1-21-1- 10011 and the University of California, San Francisco, Program for Breakthrough Biomedical Research, which is partially funded by the Sandler Foundation, to D.L.M., National Science Foundation (NSF)-1822025 to L.A.O, National Institutes of Health- National Institute of General Medical Sciences R01-GM117263-05 to J.D.B., an American Heart Association postdoctoral fellowship to F.A.-A, an NSF GRFP (DGE-1656518) and Howard Hughes Medical Institute Gilliam Fellowship (GT13330) to A.A.-B., and a DoD National Defense Science and Engineering Graduate (NDSEG) Fellowship to H.S.H. H.S.H. is a Center for Molecular Analysis and Design Fellow at Stanford University. GM/CA-XSD has been funded by the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006, P30GM138396). This research used resources of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science user facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
The U.S. Department of Energy's APS at Argonne National Laboratory is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
