A plant infected by a pathogen activates a series of biomolecular events that bestow the plant with the ability to fight off a wide range of infections, a process called systemic acquired resistance (SAR). Infection causes the plant to accumulate salicylic acid, a hormone known to activate NPR1, which is a master regulator of plant immunity. It affects more than two thousand genes involved in plant immunity and is necessary to conferring disease resistance in plants. Understanding the inner workings of NPR1 would enable major advances in our ability to increase plant immunity, but the protein is notoriously difficult to purify in a uniform conformation for structural studies, impeding in-depth understanding of its molecular biology function and regulation. By combining cryo-EM with x-ray crystallography at the U.S. Department of Energy’s Advanced Photon Source (APS), a team of researchers has uncovered the molecular basis by which salicylic acid causes NPR1 to change its shape, relocate to the nucleus, and engage with various a transcription factor to modulate expression of the plant’s defense genes. Their research, which was published in Nature, indicates new avenues for engineering disease resistance in crops while explaining decades of previous research on NPR1.
Cryo-EM of NPR1 revealed a beautiful image similar to a gliding bird with outstretched wings. As breathtaking as it was, the image did not provide sufficient resolution in areas that the team suspected were critical for the functionality of NPR1, such as densities around the protein backbone. However, the image did provide enough information to enable the scientists to combine their knowledge of protein boundary and structured regions to create a new construct that allowed them to capture the crystal structure.
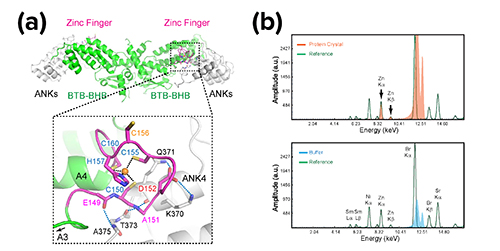
X-ray fluorescence and diffraction datasets of the crystal structure obtained at the Northeastern Collaborative Access Team (NECAT) 24-ID-C beamline at the APS (Fig. 1) revealed two surprising factors. The first was a zinc finger that wasn’t obvious from sequence analysis. (The team also used the APS with metal scan to validate the zinc.) The scientists found that not only does the zinc finger solidify the structure of the protein, but it also interfaces with a region―known as the ankyrin repeats―that interacts with the transcription factors, such as TGAs, to regulate plant immune response, thus explaining the role of the zinc finger in NPR1’s function. Importantly, these zinc-binding cysteine residues have long been shown to be critical for the NPR1 function, but how they impact function was not clear until the structural revelation. In addition to the zinc finger, the C-terminal salicylic-acid-binding domain was invisible in cryo-EM in the absent of the ligand. Biological small-angle x-ray scattering (SAXS) measurements were carried out at the APS X-ray Science Division Chemical & Materials Science Group’s beamline 12-ID-B (the APS is an Office of Science user facility at Argonne National Laboratory) and showed that the apo-form C-terminal salicylic-acid-binding domain is disordered.
Another surprise in this structural study was the palmitate, a fatty acid, that was very clearly defined in the crystal structure of the NPR1-interacting domain (NID) of TGA3. The finding is important because scientists have long conjectured a role for fatty acids in regulating the plant immune response, but the connection between the two remained unsolved. The new finding is the first to show that fatty acids are directly involved in the master immune regulator mediated transcriptional activation process.
Taken together, the findings suggest important advances in both basic research and agricultural applications. As NPR1 has such a central role in plant immunity that people have been collecting and analyzing biological data on NPR1 for decades. The new findings provide a molecular explanation for everything from genetic observations to structural analyses. The findings also suggest a new direction for people to reexamine the old literature on fatty acids and study how they can be regulated to contribute to plant immunity.
For the team, this study will serve as a starting point to build further complex systems involving the function of NPR1 as a master regulator and its role in orchestrating plants’ defenses. They will be surveying the protein’s response to a wide range of stressors through transcription activators, suppressors, and lipid metabolites that assemble on the enhancer region of a gene. ― Judy Myers
See: Shivesh Kumar1, Raul Zavaliev2, Qinglin Wu1, Ye Zhou2, Jie Cheng1, Lucas Dillard3, Jordan Powers2, John Withers2, Jinshi Zhao1, Ziqiang Guan1, Mario J. Borgnia3, Alberto Bartesaghi1,2, Xinnian Dong2*, and Pei Zhou1**, “Structural basis of NPR1 in activating plant immunity,” Nature 605, 561 (2022). DOI: 10.1038/s41586-022-04699-w
Author affiliations: 1Duke University School of Medicine, 2Duke University, 3Department of Health and Human Services
Correspondence: * xdong@duke.edu, ** peizhou@biochem.duke.edu
This work was supported by grants from the National Institutes of Health (NIH) R01 GM115355 to P.Z., R35 GM118036 to X.D., R01 GM141223 to A.B., R01 AI148366 to Z.G.; the Duke University School of Medicine Bridge Fund to P.Z.; the Howard Hughes Medical Institute to X.D.; and the Intramural Research Program of the National Institute of Environmental Health Sciences (NIEHS) grant ZIC ES103326 to M.J.B. The Northeastern Collaborative Access Team is funded by the National Institute of General Medical Sciences (grant P30 GM124165). The Pilatus 6M detector on the 24-ID-C beamline is funded by the NIH Office of Research Infrastructure Programs (Hight-End Instrumentation grant S10 RR029205). Cryo-EM data were collected using the Thermo Fisher Scientific Titan Krios transmission electron microscopes at the Janelia Research Campus and Duke University Shared Materials Instrumentation Facility (SMIF), and the Thermo Fisher Scientific Talos Arctica transmission electron microscopes at NIEHS and the University of North Carolina at Chapel Hill (UNC). This research used resources of the Advanced Photon Source, an Office of Science User Facility operated for the U.S. Department of Energy (DOE) Office of Science by Argonne National Laboratory.
The U.S. Department of Energy's APS at Argonne National Laboratory is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
