The process by which proteins fold and unfold is vital to most cellular functions. Proteins unexpectedly unfolding play an as-yet unexplained part in diseases such as cancer. To better understand unfolding, scientists need to determine the structure of the protein at each step of that process. Unfortunately, many techniques for imaging an unfolding protein average its structure either over time or spatially within a solution. Recent efforts by an international team of scientists leveraged modeling and genetic algorithms to resolve the spatial structure averaged by the x-ray data they obtained at the U.S. Department of Energy’s Advanced Photon Source (APS). Their results, published in the journal Structural Dynamics, demonstrate the success of this combination of methods by detailing the structures as a protein unfolds in real time, opening a new avenue for studying this crucial process.
To establish the success of this combination of techniques, the team of researchers from the University of Gothenburg and Lund University (both Sweden) worked with the protein apomyoglobin because it is small and has structural motifs, such as its helical backbone, in common with many other proteins. (Apomyoglobin is myoglobin without the heme coordination complex; it binds oxygen and is found in muscle tissue.) Another reason the team used apomyoglobin was because there have been previous studies of its folding structure using a wide range of methods, allowing them to compare and contrast their small-angle x-ray scattering results against the other methods. These previous studies indicated that unfolding apomyoglobin has three structures: native, molten globule, and unfolded.
First, the team created an apomyoglobin solution and measured its characteristics. Next, with colleagues from The University of Chicago, they used the BioCARS x-ray beamline 14-ID-B at the APS to take multiple types of x-ray data of the solution. (The APS is an Office of Science user facility at Argonne National Laboratory.) They collected data from solutions with different concentrations of the protein. They averaged all the small-angle x-ray scattering images taken at a given concentration to determine a steady-state value for that concentration. They subjected the solutions with higher protein concentrations to temperature jumps via an infrared laser pulse, measuring time-resolved x-ray scattering data at set time intervals following the pulse to probe protein structures at multiple length scales. To determine a reference signal value, the team also imaged the protein solution prior to the laser pulse.
To associate the time-resolved x-ray solution scattering results with physical structures of the protein, the team had to generate models of the solution where the protein existed simultaneously in multiple structural forms. They began by modeling the dynamics of a single copy of the protein, showing how its structure can change with time. They used these dynamical models to generate a large collection of possible protein structures. Then the team composed ensembles from the different structures and applied a genetic algorithm (an iterative process for optimizing the initial ensembles) to determine which ensembles produced scattering data in agreement with the measured data. The team sorted these ensembles according to structural properties, such as radius of gyration and helicity.
The team drew a number of conclusions from the measured data as well as the associations they derived between modeled ensembles and measured scattering profiles.
From the change in the measured scattering imagery as a function of time, they concluded that the protein's radius of gyration increased following the laser pulse, implying that they were observing the protein unfolding. They used singular value decomposition of the measured data to determine that three forms of the protein were in solution.
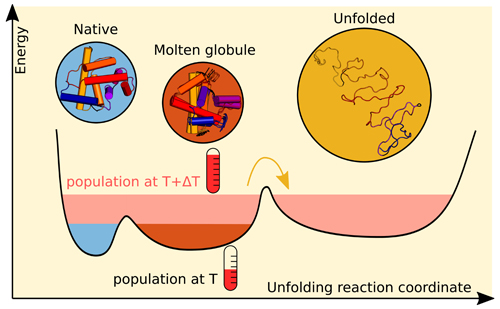
Taken together with the clusters found in the modeled structures, the team concluded these three forms were the majority in the solution at different times relative to the temperature jump, as depicted in Fig. 1. The native structure and an intermediate form were present in solution prior to application of the laser pulse, while the unfolded structure appeared approximately 1 microsecond after the temperature jump. Unlike the native and unfolded structures, the intermediate form detected by the team had different characteristics than those found by previous studies for the molten globule.
Since the previous studies were done at equilibrium and different pH levels, the team concluded that the method demonstrated in their study provides structural information for unfolding at non-equilibrium.
Taken together, the team's results demonstrate a combination of techniques for catching a protein in the act of unfolding, bringing science one step closer to understanding the unfolding process. — Mary Alexandra Agner
See: L. Henry1, M. R. Panman1, L. Isaksson1, E. Claesson1, I. Kosheleva2, R. Henning2, S. Westenhoff1*, and O. Berntsson1, 3**, “Real-time tracking of protein unfolding with time-resolved x-ray solution scattering,” Struct. Dyn. 7, 054702 (2020). DOI: 10.1063/4.0000013
Author affiliations: 1University of Gothenburg, 2The University of Chicago, 3Lund University
Correspondence: * westenho@chem.gu.se, ** oskar.berntsson@maxiv.lu.se
This research used resources of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by the Argonne National Laboratory under Contract No. DE-AC02–06CH11357. The use of BioCARS was also supported by the National Institute of General Medical Sciences of the National Institutes of Health under Grant No. P41 GM118217. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. The time-resolved setup at Sector 14 was funded in part through a collaboration with Philip Anfinrud (NIH/NIDDK). The thermostated capillary holder was developed by the group of Lin X. Chen, and we are grateful for their support when setting it up. The simulations were performed on resources provided by the Swedish National Infrastructure for Computing (SNIC) at C3SE.
The U.S. Department of Energy's APS is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
