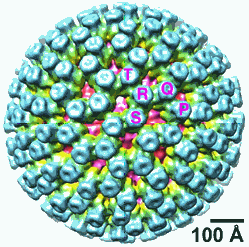
The first detailed molecular snapshots of a deadly gastrointestinal virus caught in the grasp of an immune system molecule with the capacity to inactivate it could help scientists design a more effective vaccine against rotavirus, which kills more than 500,000 children worldwide each year. The discovery was made by researchers from the Howard Hughes Medical Institute (HHMI), Harvard Medical School, and Children’s Hospital Boston using the U.S. Department of Energy’s Advanced Photon Source (APS) at Argonne National Laboratory. Their results were published in Science
Recently, the World Health Organization recommended that rotavirus vaccination be included in all national immunization programs worldwide. Virtually every child in the world becomes infected with rotaviruses before developing natural immunity. But each year, an estimated two million children are hospitalized because rotavirus infection results in severe dehydration caused by diarrhea and vomiting.
Both natural and vaccine-induced immunity occur only after the immune system has “seen” the virus and generates neutralizing antibodies. These soldiers of the immune system bind rotavirus particles, rendering them unable to infect cells.
In the new experiments, HHMI researchers mapped the structure of an antiviral antibody clamped onto a protein called VP7 that forms the rippled surface of rotavirus. The structural map reveals intimate new details about how the antibody interferes with structural changes in VP7 that help the virus infect cells. The information may be useful in designing a new generation of rotavirus vaccines that could be easier to store and administer than current vaccines, said the researchers.
Rotaviruses replicate mainly in the gut, where they infect cells in the small intestine. The virus has a triple-layered protein coat, which allows it to resist being chewed up by digestive enzymes or the gut’s acidic environment. Rotavirus does not have an envelope covering its protein shell. A virus’ envelope helps it enter host cells, and viruses without envelopes face significant hurdles in penetrating the membrane of the cells they infect. Because they have no membrane of their own, they must therefore perforate a cellular membrane to gain access to the cytoplasm (the interior of the cell.)
The new research shows that as rotavirus matures inside an infected cell, it assembles a kind of “armor” coating made of VP7 and a “spike” protein called VP4. When the mature virus particle exits one cell to infect a new cell, it perforates the endosomal membrane of the target cell by thrusting in its VP4 spike like a grappling hook.
The virus’ ability to infect cells depends on a critical structural change that quickly removes the coat of interconnected VP7 proteins—an event that unleashes the spike protein. Although researchers still do not know precisely what triggers the uncoating of VP7 during cell entry, they do know that it appears to happen when the virus enters an environment with a low concentration of calcium.
“VP7 sort of closes over VP4, locking it in place like the metal grills that surround a tree planted on a city sidewalk,” explained HHMI investigator Stephen C. Harrison, a co-author on the paper. “And it is the loss of VP7 in the uncoating step that triggers VP4 to carry out its task.”
To get a closer look at how antibodies latch onto VP7 and neutralize the virus, Harrison and his colleagues from Children’s Hospital, Stanford University School of Medicine, VA Palo Alto Health Care System, and HHMI used x-ray crystallography at the Northeastern Collaborative Access Team 24-ID-C beamline at the APS to examine the molecular architecture of VP7 in the grasp of a fragment of the antibody. X-ray crystallography is a powerful tool for “seeing” the orientation of atoms and the distances separating them within the molecules.
Before Harrison’s team could use x-ray crystallography, however, they first had to crystallize VP7 in complex with the antibody fragment. Only after that step was completed could they move on to bombarding those crystallized proteins with x-rays. Computers helped capture the diffraction patterns that emerged as the x-rays scattered from the crystal lattice. By rotating the crystallized protein complexes through multiple exposures, the researchers could record enough data to calculate three-dimensional models, which exposed the underlying architecture of VP7 and the antibody fragment.
The resulting detailed structural map of the VP7-antibody protein complex revealed that the antibody neutralizes the virus by preventing the VP7 proteins from dissociating. Normally, calcium creates a bridge between VP7 molecules that holds them in place until uncoating. This structure revealed that the antibody makes an additional bridge, cementing the subunits together, making the virus resistant to the uncoating trigger and preventing it from infecting cells.
Current rotavirus vaccines consist of weakened live virus that triggers the immune system to produce neutralizing antibodies. However, the new structural findings suggest how researchers might engineer a different type of rotavirus vaccine consisting only of immune-triggering protein, said Harrison. This protein-only vaccine could be made of a chemically linked complex of VP7 molecules that would stimulate the immune system to produce antibodies that block rotavirus infection more effectively than antibodies made in response to immunization with VP7 that is not cross-linked.
While live-virus-based vaccines have been effective, they have drawbacks that a protein-based vaccine might overcome. The virus-based vaccines are perishable and require refrigeration, but vaccines based on proteins could be more stable and stored at room temperature. Another benefit is that protein-based vaccines could be combined with other protein vaccines in a “cocktail” that would cut down on the number of immunizations since blending cannot be done so readily with virus-based vaccines. These advantages could make protein vaccines especially useful in developing countries that lack an extensive public health infrastructure and where the vast majority of childhood deaths from rotavirus occur.
Contacts: S.C. Harrison harrison@crystal.harvard.edu,
P.R. Dormizer philip.dormitzer@novartis.com
See: Scott T. Aoki, Ethan C. Settembre, Shane D. Trask, Harry B. Greenberg, Stephen C. Harrison, and Philip R. Dormitzer, “Structure of Rotavirus Outer-Layer Protein VP7 Bound with a Neutralizing Fab,” Science, 324(5933),1444 (12 June 2009). DOI: 10.1126/science.1170481
Supported by National Institutes of Health (NIH) grant CA-13202 (S.C.H.), by a VA Merit Award and NIH grants AI-21362 and DK-56339 (H.B.G.), and by an Ellison Medical Foundation New Investigators in Infectious Diseases Award (P.R.D.). S.C.H. is an Investigator of the HHMI.
Use of the Advanced Photon Source at Argonne National Laboratory was supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences (DOE-BES), under Contract No. DE-AC02-06CH11357.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America 's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC for the U.S. Department of Energy's Office of Science.
The original HHMI news release can be found at:
http://www.hhmi.org/news/new-images-may-improve-vaccine-design-deadly-rotavirus