Imagine having to copy an entire book by hand without missing a comma. Our cells face a similar task every time they divide. They must duplicate both their DNA and a subtle pattern of punctuation-like modifications on the DNA known as methylation.
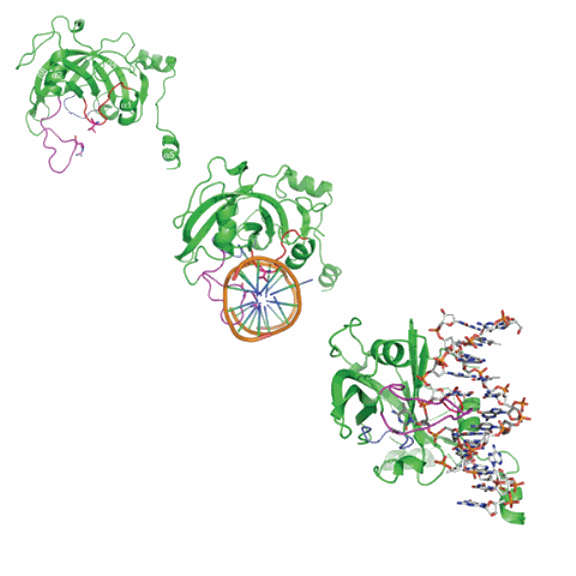
Scientists at Emory University School of Medicine and the University of California, Los Angeles, have caught in action one of the tools mammalian cells use to maintain their pattern of methylation. Visualized by x-ray crystallography carried out at the Southeast Regional Collaborative Access Team (SER-CAT) beamline 22-ID-D at the U.S. Department of Energy’s Argonne Advanced Photon Source (SER-CAT), the SRA domain of the protein UHRF1 appears to act like a bookmark while enzymes are copying a molecule of DNA.
The team's description of the protein's structure while bound to DNA was published in Nature.
Scientists refer to methylation, the addition of a methyl group to DNA, as an "epigenetic" modification because it adds a layer of information on top of the genetic sequence of the DNA itself. It marks genes for silencing, which means they do not manufacture proteins.
"The processes that copy the methylation pattern have to be faithful," says senior author Xiaodong Cheng, professor of biochemistry at Emory. "Otherwise, losing DNA methylation marks can have serious consequences, causing genes to become active at the wrong places and times."
"Gene silencing via DNA methylation is critical for normal development and for curbing the runaway cell division that characterizes cancer," said Peter Preusch, who oversees biophysics grants at the National Institute of General Medical Sciences of the National Institutes of Health. "Alterations in methylation patterns are also important for generating embryonic stem-like cells from differentiated cells."
In mammalian cells, methylation usually appears on double-stranded DNA where the nucleotide Cytosine (C) is followed by Guanine (G). The complementary sequence on the opposite strand is also C then G, and the methylation appears on both Cs.
When a cell is copying its DNA, a set of enzymes duplicates the DNA sequence from the parental strand to the new "daughter" strand but not the methylation. Each new daughter strand of the DNA molecule is left with the previously methylated Cs unmethylated. UHRF1 recognizes this "hemi-methylated" DNA and calls in a methyltransferase enzyme to add a second methyl group onto the daughter strand.
"UHRF1 has the important task of making sure the methyltransferase enzyme does its job in the right place and right time," Cheng says.
Mouse cells that have deleted the UHRF1 gene are more sensitive to DNA-damaging agents such as radiation, and mouse embryos without the gene cannot complete development. Other studies have found that cancer cells produce more UHRF1 than non-cancerous cells.
What was an unexpected finding was how the SRA domain of UHRF1 recognizes the hemi-methylated DNA, Cheng says. It flips the methylated nucleotide out of the DNA helix, which only had been seen previously in enzymes that physically modify the DNA.
Cheng says the flipping mechanism could prevent the protein from sliding away once it has found a hemi-methylated site. "It suggests that it serves as a placeholder, where it recruits other enzymes for faithful DNA methylation or repair enzymes if the DNA has been damaged," he says.
Contact: Xiaodong Cheng, xcheng@emory.edu
See: Hideharu Hashimoto, John R. Horton, Xing Zhang, Magnolia Bostick, Steven E. Jacobsen, and Xiaodong Cheng; "The SRA domain of UHRF1 flips 5-methylcytosine out of the DNA helix," Nature advance online publication 3 September 2008 | doi:10.1038/nature07280
Press release source: Emory University
The research was funded by the National Institutes of Health, the Howard Hughes Medical Institute and the Georgia Research Alliance. SER-CAT is funded mainly through state legislative funds, agencies, and the individual universities at the university, department, or individual research group levels, and is operated by the University of Georgia. The Advanced Photon Source at Argonne National Laboratory is supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357.