BY ANDRE SALLES | MARCH 2, 2023
Scientists at the Advanced Photon Source are exploring ways to analyze X-ray data faster and with more precision. One new software package called TomocuPy has shown to be up to 30 times faster than the current practice.
When it comes to X-ray data collection and processing, one goal remains constant: do it faster.
This is especially important for scientists working at the Advanced Photon Source (APS), a U.S. Department of Energy (DOE) Office of Science user facility at DOE’s Argonne National Laboratory. The APS is about to undergo an extensive upgrade, one that will increase the brightness of its X-ray beams by up to 500 times. Scientists use those beams to see ions moving inside batteries, for instance, or to determine the exact protein structure of infectious diseases. When the upgraded APS emerges in 2024, they will be able to collect that data at an exponentially faster rate.
But to keep pace with the science, the analysis and reconstruction of that data — which shapes it into a useful form — will also have to get much faster before the APS Upgrade is complete. Data from different X-ray techniques are analyzed in different ways, but Argonne scientists have been working on multiple new methods using artificial intelligence to help speed up the timeline. Faster processes have been created for X-ray imaging and for determining important data peaks in X-ray diffraction data, to name a couple.
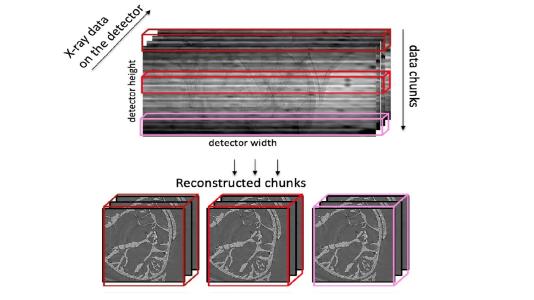
Argonne’s Viktor Nikitin, an assistant physicist working at the APS, has now unveiled a new way of reconstructing data taken through a process called tomography. Nikitin’s software package, called TomocuPy, builds on the current tools scientists use for tomography data. It improves the speed of the process by 20 to 30 times by leveraging computers equipped with graphics processing units (GPUs) and by reconstructing several chunks of data at once. Nikitin is the sole author of a paper explaining TomocuPy, published in the Journal of Synchrotron Radiation, though he’s quick to note that he didn’t work in a vacuum: he had help from his APS colleagues (including his group leader, Francesco De Carlo) and he built on previous data reconstruction methods developed to run on central processing units (CPUs).
But Nikitin’s innovations are important, and they involve an understanding of how tomography works: slice by slice. Tomography involves using an X-ray beam to observe multiple parts of the sample, extracting cross sections (or slices) from them, and then using a computer to reconstruct those slices into a whole. Nikitin’s TomocuPy builds a pipeline for processing those slices where the sequence of operations, such as reading and writing from hard disks and computations, can happen concurrently. Current methods examine each slice one at a time and put them together on the back end.
TomocuPy also takes advantage of the multiple processors within each GPU being used, and runs them all simultaneously. Stack up enough of these, and you can look at thousands of slices in the time it would previously have taken to analyze one. Nikitin’s method also saves computing time by lowering the analysis precision of each GPU to match the output from the detector — if the output is 16-bit, he says, you don’t need 32-bit calculations to analyze it.
“Tomography is a lot of small operations, processing small images,” Nikitin said. “GPUs can do it up to 30 times faster. The previous method uses CPUs and doesn’t use the information pipeline that TomocuPy does, and it’s far slower.”
That pipeline includes a streaming process to move data between various applications. That process was developed by Argonne’s Siniša Veseli, a principal software engineer. The framework in use at the APS has limits on the amount of data that can be moved, but that data can now be distributed to multiple different applications at once. This, Veseli says, helps the information highway at the APS keep up with the torrent of data coming off of its X-ray detectors.
“After the APS Upgrade, computing rates are going to be much higher,” he said. “We will have new detectors and the accelerator will be at a higher brilliance. We need to come up with tools that are able to handle this scale of image rate. Now everything is done in real time over the network.”
The GPUs in use for TomocuPy are often used for artificial intelligence applications, and can be adapted to work with machine learning algorithms. This is important, Nikitin said, because the eventual goal is experiments that can adjust to reconstructed data in real time. As part of the imaging group at the APS, Nikitin sees firsthand the need for such speed. He gives an example of an experiment using pressure to test the durability of a new material.
“You apply pressure and a microscopic crack appears. As you increase the pressure, you want to zoom in on that region, but it’s always hard to find where the crack is,” he said. “We want to have the data reconstruction immediately to find the region where you want to zoom in. Before, it would take 10 to 15 minutes, and the crack will have formed already.”
Nikitin and his colleagues used a similarly dynamic experiment — tracking the formation of gas hydrates inside a porous sample at beamline 2-BM of the APS — to test TomocuPy’s efficiency. Gas hydrates are tiny particles that look like ice, but contain gases (methane, in this case) and form under cold conditions. The team used various GPU resources, including the Polaris supercomputer at the Argonne Leadership Computing Facility (ALCF), to demonstrate the tool’s ability to reconstruct X-ray data quickly and accurately. They were able to reconstruct data and use it to find areas of interest far faster than previous methods allowed. The ALCF is a DOE Office of Science user facility.
Eventually, Nikitin said, the plan is to use artificial intelligence to help direct experiments, either by automatically zooming in on the interesting parts of a sample, or changing the environmental conditions like temperature and pressure in response to quickly reconstructed huge amount of APS Upgrade data. This will be possible, he said, with help from the massive supercomputers at the ALCF.
“We are building a fast connection between APS and ALCF,” he said. “By running TomocuPy on a supercomputer, we can do in a day what now can take up to a month.”
