The original Northwestern University press release can be read here.
Scientists are already preparing for a possible next coronavirus pandemic to strike. In future-looking research, Northwestern University Feinberg School of Medicine scientists have identified a novel target for a drug to treat SARS-CoV-2 that also could impact a new emerging coronavirus.
"God forbid we need this, but we will be ready," said Karla Satchell, professor of microbiology-immunology at Feinberg, who leads an international team of scientists to analyze the important structures of the virus.
The Northwestern team previously mapped the structure of a virus protein called nsp16, which is present in all coronaviruses. This new study provides critical information that could aid drug development against future coronaviruses as well as SARS-CoV-2.
"There is great need for new approaches to drug discovery to combat the SARS-CoV-2/COVID-19 pandemic and infections from future coronaviruses," Satchell said. "The idea is this future drug would work early in the infection," Satchell said. "If somebody around you gets the coronavirus, you would run to the drugstore to get your medication and take it for three or four days. If you were sick, you wouldn't get as sick."
The paper was published in Science Signaling.
Satchell's team has mapped or "solved" three new protein structures in three-dimensional views and discovered a secret identifier in the machinery that helps the virus hide from the immune system.
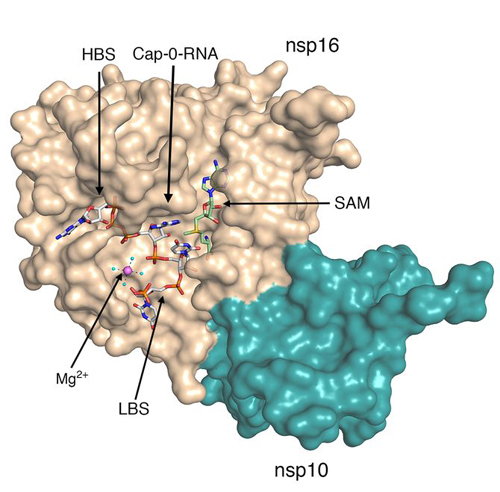
They discovered a coronavirus-specific pocket in the protein, nsp16, that binds the virus-genomic fragment held in place by a metal ion. The fragment is used by the coronavirus as the template for all the viral building blocks.
For this reason, Satchell said, there is potential to make a drug to fit this unique pocket that would block function of this protein from the coronavirus. It would not block the function of a similar protein from human cells that lacks the pocket. Thus, such a drug would only target the invader protein.
Nsp16 is considered one of the key viral proteins that could be inhibited by drugs to stop the virus shortly after a person gets exposed. The goal is to stop the virus early before people get too sick. Since little research was done on nsp16, Satchell's team has worked to generate key information about this protein and is collaborating with chemists who will use the information to design drugs against the protein.
While some of the coronavirus proteins vary a lot, nsp16 is nearly the same across most of them. The unique pocket discovered by Satchell's group is present in all the different coronavirus members. This means that drugs designed to fit this pocket should work against all coronaviruses, including a virus that emerges in the future. And it should work against the common cold that is caused by a coronavirus.
Satchell envisions any drug developed from her team's discovery of the coronavirus pocket would be part of a treatment cocktail taken by patients early in the course of the disease. That could include drugs similar to Remdesivir, a drug that prevents the virus from producing the template for the building blocks that is necessary for it to replicate itself.
The Northwestern team in the Center for Structural Genomics of Infectious Diseases (CSGID) expressed, purified and crystallized this protein. The idea of the project came from first study author George Minasov, research associate professor of microbiology-immunology at Feinberg. He worked with Feinberg research associate professor Ludmilla Shuvalova to crystallize the protein and also with post-doctoral fellow Monica Rosas-Lemus, who developed an assay to test the function of the protein based on information from the structure.
The team collaborated with Purdue University investigator Andrew Mesecar, who helped with biochemistry assays. Data on the structure was collected at the Life Sciences Collaborative Access Team [beamlines 21-ID-D and 21-ID-F at the Advanced Photon Source, a Department of Energy Office of Science user facility at Argonne National Laboratory] by Joseph Brunzelle. Minasov solved the structure from the collected data. This project is one of many carried out by the CSGID to use structural biology to understand the biology of the virus responsible for the COVID-19 pandemic.
Overall, the center has made significant contributions to the development of vaccines, drugs and diagnostics. The international team has solved more than 70 different viral structures to reveal viral protein structure, interactions with possible drugs and interactions with antibodies. This work is made freely available to the global community to use to accelerate efforts to design new treatments against coronavirus to combat COVID-19 and future pandemics.
See: George Minasov et al., "Mn2+ coordinates Cap-0-RNA to align substrates for efficient 2′-O-methyl transfer by SARS-CoV-2 nsp16," Sci. Signal.
10.1126/scisignal.abh2071 (2021).
Correspondence: k-satchell@northwestern.edu
Use of the Life Sciences Collaborative Access Team was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor (Grant 085P1000817). Funding: NIAID/NIH/HHS Contract HHSN272201700060C (A.M. and K.S.); Chicago Biomedical Consortium COVID Response Award CR-003 (K.S.); NCI CCSG P30 CA060553 (K.S. and HTAL core facility); P.H. was funded by NCI R01-CA142861 (to L. Laimins). This research used resources of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract DE-AC02-06CH11357.
Image of nsp16 from George Minasov et al., "Mn2+ coordinates Cap-0-RNA to align substrates for efficient 2′-O-methyl transfer by SARS-CoV-2 nsp16," Sci. Signal. 10.1126/scisignal.abh2071 (2021). © 2021 American Association for the Advancement of Science. All rights reserved.
The U.S. Department of Energy's APS is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
